2 Institute of Life Sciences, Jiyang Colloge of Zhejiang A&F University, Zhuji, 311800, Zhejiang, China
 Author
Author  Correspondence author
Correspondence author
International Journal of Super Species Research, 2024, Vol. 14, No. 3
Received: 21 Apr., 2024 Accepted: 23 May, 2024 Published: 30 May, 2024
Human Immunodeficiency Virus (HIV) remains a formidable challenge due to its remarkable ability to adapt and evade both the immune system and antiretroviral therapies. This feature review delves into the multifaceted strategies employed by HIV to ensure its persistence and proliferation. Central to HIV's adaptability is its genetic diversity, driven by high mutation rates and recombination events duringreplication. These mechanisms enable the virus to rapidly develop resistance to antiretroviral drugs and escape immune responses, complicating efforts to design effective vaccines and treatments. The virus's ability to form a latent reservoir in resting memory CD4+T cells further complicates eradication efforts, as these reservoirs harbor escape mutations that render infected cells insensitive to cytotoxic T lymphocytes. Additionally, the continuous evolution of new HIV subtypes poses significant challenges for disease management and vaccine development. Understanding these adaptation strategies is crucial for developing innovative therapeutic approaches and ultimately achieving a cure for HIV.
1 Introduction
Human Immunodeficiency Virus (HIV) remains a formidable global health challenge despite significant advancements in medical research and treatment. Since its identification, HIV has infected millions of individuals worldwide, leading to the Acquired Immunodeficiency Syndrome (AIDS) and causing substantial morbidity and mortality. The virus's ability to evade the immune system and develop resistance to antiretroviral therapies (ART) complicates efforts to control its spread and achieve a cure (Ávila-Ríos et al., 2019; Cabrera-Rodríguez et al., 2021; Ta et al., 2022). The persistence of HIV in latent reservoirs within the body necessitates lifelong treatment for those infected, posing ongoing physiological and economic burdens (Ta et al., 2022).
Understanding the adaptation strategies of HIV is crucial for developing effective treatment and prevention methods. HIV's ability to adapt to host immune pressures, particularly through interactions with human leukocyte antigen (HLA) molecules, significantly impacts the virus's evolution and the clinical outcomes of infected individuals (Carlson et al., 2016; Ávila-Ríos et al., 2019). These adaptations can lead to immune escape variants that complicate vaccine development and hinder the effectiveness of immune responses (Carlson et al., 2016; Ávila-Ríos et al., 2019). Additionally, the interplay between HIV and cellular processes such as autophagy highlights potential targets for therapeutic intervention (Cabrera-Rodríguez et al., 2021). By elucidating these adaptation mechanisms, researchers can identify novel strategies to enhance immune control, design more effective vaccines, and develop approaches to eradicate latent viral reservoirs (Ávila-Ríos et al., 2019; Cabrera-Rodríguez et al., 2021; Ta et al., 2022).
This study provides a comprehensive overview of the current understanding of HIV's adaptation strategies and their implications for treatment and prevention. Researchers explore the clinical and evolutionary consequences of HIV adaptation to HLA-associated immune pressures, the impact of pre-adapted HIV transmission, and the role of autophagy in early HIV infection. Additionally, researchers will discuss the potential of digital health interventions and novel therapeutic strategies to address the evolving landscape of HIV treatment and prevention. By synthesizing recent findings, this review seeks to inform future research directions and contribute to the ongoing efforts to combat HIV/AIDS effectively.
2 Biology of HIV
2.1 Structure and Genetic Makeup of the HIV Virus
Human Immunodeficiency Virus (HIV) is a retrovirus that primarily targets the human immune system, specifically CD4+T cells, macrophages, and dendritic cells. The virus is composed of two main types, HIV-1 and HIV-2, each with distinct genetic and structural characteristics. HIV-1 is the more prevalent and virulent form globally, while HIV-2 is less transmissible and largely confined to West Africa (Calado et al., 2023).
The HIV virion is spherical and enveloped, containing two copies of single-stranded RNA. The viral envelope is derived from the host cell membrane and is embedded with glycoproteins, primarily gp120 and gp41, which are crucial for the virus's ability to bind and enter host cells (Mcdougal et al., 1986). Inside the envelope, the virus has a conical capsid made of p24 protein, which encases the RNA genome and essential enzymes such as reverse transcriptase, integrase, and protease (Azimi and Lee, 2020). These enzymes are vital for the virus's replication cycle, allowing it to reverse transcribe its RNA into DNA, integrate into the host genome, and produce new viral particles.
2.2 HIV Life Cycle: From Entry into the Host Cell to Replication
The HIV life cycle begins with the virus binding to the CD4 receptor on the surface of a target cell, typically a CD4+T cell, macrophage, or dendritic cell. This binding is mediated by the viral glycoprotein gp120, which also interacts with a co-receptor, either CCR5 or CXCR4, facilitating the fusion of the viral envelope with the host cell membrane (Poli, 2013). Once inside the cell, the viral RNA is reverse transcribed into DNA by the enzyme reverse transcriptase (Yin et al., 2020).
The newly synthesized viral DNA is then transported into the nucleus, where it integrates into the host cell's genome with the help of the integrase enzyme. This integrated viral DNA, known as a provirus, can remain latent or be transcribed into RNA, which serves as the genome for new virions and as mRNA for the production of viral proteins (Yin et al., 2020). The viral proteins and RNA are assembled into new virions, which bud off from the host cell, acquiring a portion of the host cell membrane as their envelope. This process is facilitated by the protease enzyme, which cleaves precursor proteins into their functional forms (Azimi and Lee, 2020).
2.3 Types of HIV: HIV-1 vs. HIV-2 and Their Global Distribution
HIV-1 and HIV-2 are the two main types of HIV, each with distinct epidemiological and clinical characteristics. HIV-1 is the most widespread and is responsible for the global HIV pandemic. It is highly transmissible and progresses more rapidly to AIDS compared to HIV-2. HIV-1 is further classified into several groups (M, N, O, and P), with Group M being the most prevalent and subdivided into multiple subtypes (A, B, C, etc.) (Calado et al., 2023).
HIV-2, on the other hand, is less transmissible and is primarily found in West Africa. It has a longer asymptomatic period and a slower progression to AIDS. The lower virulence of HIV-2 is partly due to its reduced ability to replicate in host cells and its less efficient transmission routes (Calado et al., 2023) Despite these differences, both types of HIV share similar mechanisms of immune evasion and pathogenesis, including the ability to infect and manipulate dendritic cells to enhance their spread and persistence within the host (Cameron et al., 1992; Tsunetsugu-Yokota et al., 2013; Martín-Moreno et al., 2019).
Understanding the biology of HIV, including its structure, life cycle, and the differences between HIV-1 and HIV-2, is crucial for developing effective treatments and vaccines. The virus's ability to adapt and evade the immune system poses significant challenges, but ongoing research continues to uncover new insights into its complex interactions with the host immune system (Bayliss et al., 2020; Mulherkar et al., 2022).
3 Mechanisms of HIV Adaptation
3.1 High Mutation Rate and Genetic Variability
One of the most significant factors contributing to HIV's adaptability is its exceptionally high mutation rate. HIV-1 reverse transcriptase (RT), the enzyme responsible for converting viral RNA into DNA, is notably error-prone due to its lack of proofreading capabilities. This low fidelity results in a high rate of mutagenesis, which is a key factor in the virus's ability to escape immune responses and develop resistance to antiretroviral drugs (Figure 1) (Bowen et al., 2022).
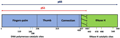 Figure 1 Schematic representation of HIV-1 RT structure (Adopted from Bowen et al., 2022) Image caption: HIV-1 RT consists of two subunits, p66 and p51. The p66 subunit harbors both DNA polymerase (blue segment) and RNase H (green segment) domains, whereas p51, which is a proteolytic cleavage product of p66 subunit, does not have an active RNase H domain (residue 427–440). HIV-1 RT DNA polymerase domain, which captures DNA primer annealed to RNA or DNA template, in the form of a clasping right hand, can be further classified into the finger, palm, and thumb subdomains, whereas the connection subdomain links polymerase and RNase H domains. Amino-acid residues crucial for the enzymatic catalytic sites of DNA polymerase and RNase H activities are marked with respective colors (Adopted from Bowen et al., 2022) |
Studies have shown that the mutation rate of HIV-1 in vivo is extraordinarily high, with estimates of (4.1±1.7)×10⁻3 per base per cell, the highest reported for any biological entity. This high mutation rate is not solely due to the intrinsic properties of RT but is also significantly influenced by host factors such as cytidine deaminases of the A3 family, which contribute to the majority of mutations (Cuevas et al., 2015). These mutations can lead to the generation of drug-resistant and immunological-escape mutants, complicating treatment and vaccine development efforts (Abram et al., 2014).
3.2 Antigenic Variation: Evasion of the Host Immune Response
HIV's ability to evade the host immune system is another critical adaptation strategy. The virus achieves this through antigenic variation, which involves frequent changes in the viral envelope proteins that are the primary targets of the host's immune response. This constant variation makes it difficult for the immune system to recognize and neutralize the virus effectively.
The high mutation rate of HIV-1 RT plays a crucial role in this process by generating a diverse population of viral variants. These variants can present different antigens to the immune system, allowing some to escape immune detection and continue replicating. This mechanism is particularly effective in evading neutralizing antibodies, which are a key component of the adaptive immune response (Cuevas et al., 2015; Bowen et al., 2022).
3.3 Integration into the Host Genome and Establishment of Viral Reservoirs
A defining feature of HIV is its ability to integrate its genetic material into the host genome, establishing long-term viral reservoirs. After the viral RNA is reverse-transcribed into DNA by RT, the resulting proviral DNA is integrated into the host cell's genome. This integration is a critical step in the viral life cycle and is essential for the persistence of the virus within the host (Bowen et al., 2022).
Once integrated, the proviral DNA can remain latent for extended periods, evading detection by the immune system and antiretroviral drugs. These latent reservoirs are primarily found in resting memory CD4+T cells and other long-lived cells, making them a significant barrier to curing HIV infection. The establishment of these reservoirs ensures that the virus can persist even in the face of potent antiretroviral therapy, leading to viral rebound if treatment is interrupted (Bowen et al., 2022).
In conclusion, HIV's adaptation strategies, including its high mutation rate, antigenic variation, and integration into the host genome, underscore the complexity of combating this virus. These mechanisms not only facilitate the virus's evasion of the immune system but also contribute to the challenges in developing effective treatments and vaccines. Understanding these strategies in greater detail is crucial for advancing HIV research and improving therapeutic interventions.
4 Immune Evasion Strategies
4.1 Downregulation of Host Immune Receptors (e.g., CD4, MHC-I)
HIV has developed sophisticated mechanisms to downregulate key immune receptors, such as CD4 and MHC-I, to evade the host immune system. The downregulation of CD4, the primary receptor for HIV entry, is crucial for preserving viral infectivity. This process involves three HIV-1 proteins: Nef, Env, and Vpu. Env forms a complex with CD4 in the endoplasmic reticulum, retaining the receptor in this compartment. Nef and Vpu act as connectors between CD4 and specific intracellular trafficking pathways, targeting the receptor for degradation in the lysosome and the proteasome, respectively. This downregulation not only prevents superinfection but also reduces the visibility of infected cells to the immune system.
Similarly, the downregulation of MHC-I, primarily mediated by the Nef protein, involves the redistribution of this receptor, leading to its accumulation in the Golgi apparatus. This process is less understood but is believed to play a significant role in immune evasion by preventing the presentation of viral peptides to cytotoxic T lymphocytes (CTLs). The ability of HIV to modulate these receptors highlights its capacity to interfere with the host's immune surveillance mechanisms, thereby enhancing its survival and persistence.
4.2 Latency and the Hidden Reservoirs: Avoidance of Immune Detection
One of the most formidable challenges in combating HIV is its ability to establish latent reservoirs. These reservoirs consist of infected CD4+T cells that harbor replication-competent virus but remain transcriptionally silent, evading detection by the immune system and antiretroviral therapy (ART). The "kick-and-kill" strategy, which involves reactivating latent virus followed by immune-mediated clearance, has shown promise in latency models but has not been fully effective in clinical settings. This is partly due to the inherent resistance of latently infected cells to CD8+ T cell-mediated killing (Huang et al., 2018).
The establishment of latency is closely linked to the transition of CD4+ T cells from an effector to a memory state. During this transition, there is a temporary upregulation of CCR5 expression and a rapid downregulation of cellular gene transcription, creating a window of opportunity for HIV to integrate into the host genome and establish latency. This process is further complicated by the presence of a reservoir-enriched lymphoid microenvironment, which resembles the tumor microenvironment and limits the access and function of CD8+T cells (Shan et al., 2017). Understanding the mechanisms underlying latency and developing strategies to target these hidden reservoirs are critical for achieving a functional cure for HIV (Figure 2) (Delannoy et al., 2019; Mylvaganam et al., 2019).
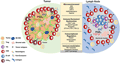 Figure 2 Parallels between immunoregulation in solid tumors and lymph nodes (Adopted from Mylvaganam et al., 2019) Image caption: Tumors and LNs are composed of stromal and immune cells that secrete cytokines and growth factors such as transforming growth factor b (TGFβ), prostaglandin E2 (PGE2), indolamine 2-3-dioxygenase (IDO), and adenosine that shape the tumor and LN microenvironment and collectively contribute to suppression of the T cell response. Adenosine signals through the adenosine 2A receptor (A2AR) and promotes production of cyclic AMP, which impairs T cell trafficking, proliferation, and cytotoxicity (73). Immunosuppression is also induced by regulatory T cells (Treg) that express higher levels of CTLA-4, an inhibitory receptor that outcompetes CD80/CD86 on the surface of effector cells and promotes the production of IDO, an enzyme that degrades tryptophan and leads to impaired proliferation and Treg differentiation. Additionally, Tregs express high levels of CD73/CD39, enzymes that convert ATP to adenosine, which inhibit immune function; Tumors upregulate inhibitory ligands such as PD-L1 that bind to inhibitory receptors resulting in suppression of adaptive immune responses; Several strategies have been developed in the immune-oncology field to overcome these barriers such as checkpoint blockade, small molecule inhibitors, therapeutic vaccines, and CAR T cell therapy. These immune based therapies can all be extended to the HIV cure field. Several mechanisms that result in resistance to checkpoint blockade and CD8 T cell exclusion include activation of the WNT/β -catenin pathway, localization of M2 macrophages within the tumor, and the secretion of TGFβ (Adopted from Mylvaganam et al., 2019) |
4.3 Impact on Host Immune Cells: CD4 T Cell Depletion and Immune Dysfunction
HIV infection leads to a dramatic depletion of CD4+T cells, which are crucial for orchestrating the immune response. This depletion is a hallmark of HIV-induced immune dysfunction and is primarily driven by direct viral killing of infected cells, immune-mediated killing, and chronic immune activation. The loss of CD4+T cells impairs the host's ability to mount effective immune responses against opportunistic infections and malignancies, contributing to the progression to AIDS (Mylvaganam et al., 2019).
Moreover, HIV-induced immune dysfunction extends beyond CD4+ T cell depletion. The virus also disrupts the function of surviving immune cells. For instance, HIV-1 Nef-mediated downregulation of MHC-I on infected cells impairs the ability of CTLs to recognize and eliminate these cells, further compromising the immune response (Mujib et al., 2017). Additionally, the chronic activation of the immune system in response to ongoing viral replication leads to immune exhaustion, characterized by the upregulation of inhibitory receptors such as PD-1 on T cells. This state of exhaustion reduces the efficacy of immune responses and is a significant barrier to achieving viral control (Mylvaganam et al., 2019).
In summary, HIV employs multiple strategies to evade the host immune system, including the downregulation of key immune receptors, the establishment of latent reservoirs, and the induction of immune dysfunction. These mechanisms not only facilitate viral persistence but also complicate efforts to develop effective therapies and achieve a cure. Understanding these evasion strategies is essential for designing interventions that can overcome these barriers and ultimately eradicate HIV.
4.3 Impact on Host Immune Cells: CD4 T Cell Depletion and Immune Dysfunction
HIV infection leads to a dramatic depletion of CD4+T cells, which are crucial for orchestrating the immune response. This depletion is a hallmark of HIV-induced immune dysfunction and is primarily driven by direct viral killing of infected cells, immune-mediated killing, and chronic immune activation. The loss of CD4+T cells impairs the host's ability to mount effective immune responses against opportunistic infections and malignancies, contributing to the progression to AIDS (Mylvaganam et al., 2019).
Moreover, HIV-induced immune dysfunction extends beyond CD4+ T cell depletion. The virus also disrupts the function of surviving immune cells. For instance, HIV-1 Nef-mediated downregulation of MHC-I on infected cells impairs the ability of CTLs to recognize and eliminate these cells, further compromising the immune response (Mujib et al., 2017). Additionally, the chronic activation of the immune system in response to ongoing viral replication leads to immune exhaustion, characterized by the upregulation of inhibitory receptors such as PD-1 on T cells. This state of exhaustion reduces the efficacy of immune responses and is a significant barrier to achieving viral control (Mylvaganam et al., 2019).
In summary, HIV employs multiple strategies to evade the host immune system, including the downregulation of key immune receptors, the establishment of latent reservoirs, and the induction of immune dysfunction. These mechanisms not only facilitate viral persistence but also complicate efforts to develop effective therapies and achieve a cure. Understanding these evasion strategies is essential for designing interventions that can overcome these barriers and ultimately eradicate HIV.
5 Drug Resistance Mechanisms
5.1 Mutation-driven resistance to antiretroviral therapy (ART)
HIV's ability to rapidly mutate is a primary driver of resistance to antiretroviral therapy (ART). The virus's high replication rate and lack of repair mechanisms contribute to a high mutation frequency, which can lead to the development of drug-resistant strains (Steiner et al., 2020). Drug-resistance mutations (DRMs) often incur a fitness penalty to the virus, but these are compensated by accessory mutations that interact favorably with the viral genome, creating an evolutionary trapping effect (Choudhuri et al., 2022). This process is complex and involves multiple time scales, with some mutations becoming entrenched due to the collective effect of many accessory mutations rather than a few strongly coupled ones (Choudhuri et al., 2022). The rapid evolution of HIV under drug-selection pressure necessitates continuous monitoring and adaptation of ART regimens to manage and mitigate esistance (Clutter et al., 2016).
5.2 The role of viral diversity in the emergence of drug-resistant strains
Genetic diversity within HIV populations plays a crucial role in the emergence of drug-resistant strains. This diversity is driven by the high mutation rate during DNA synthesis by reverse transcriptase, large viral populations, rapid viral turnover, and high recombination rates (Martinez-Picado and Martínez, 2008). The presence of multiple HIV subtypes and recombinant forms further complicates the landscape of drug resistance. For instance, in Cameroon, the CRF02 (A/G) subtype was found to be the most prevalent, with significant differences in resistance mutation profiles between ART-experienced and ART-naive patients (Meriki et al., 2019). This diversity allows the virus to escape host immune responses and develop resistance to various antiretroviral drugs, making it challenging to design universally effective therapies (Martinez-Picado and Martínez, 2008).
5.3 Challenges in designing effective therapies due to rapid viral evolution
The rapid evolution of HIV poses significant challenges in designing effective therapies. The virus's ability to quickly develop resistance to ART means that treatment regimens must be continuously updated and tailored to individual patients' viral genotypes (Clutter et al., 2016). The emergence of drug-resistant mutations can lead to virological failure and the accumulation of further resistance, complicating long-term management of HIV infection (Paredes and Clotet, 2010). Additionally, the increasing complexity of antiretroviral regimens has favored the selection of HIV variants harboring multiple drug resistance mutations, which can severely impact viral fitness and treatment outcomes (Martinez-Picado and Martínez, 2008). The development of new drugs with higher genetic barriers to resistance, such as integrase strand transfer inhibitors, is crucial to counteract the rapid evolution of HIV and maintain the effectiveness of ART (Iyidogan and Anderson, 2014; Meriki et al., 2019).
In summary, understanding the mechanisms of HIV drug resistance, the role of viral diversity, and the challenges posed by rapid viral evolution is essential for developing effective treatment strategies. Continuous monitoring, personalized therapy, and the development of new drugs with higher resistance barriers are key to managing HIV in the long term.
6 Case Study: The Emergence of Multi-Drug Resistant HIV Strains
6.1 Overview of Multi-Drug Resistant HIV Cases in Specific Regions
The emergence of multi-drug resistant (MDR) HIV strains has been documented globally, with significant variations in prevalence across different regions. In high-income countries, the widespread use of antiretroviral therapy (ART) has led to a notable reduction in HIV-related mortality and incidence. However, this has also resulted in the selection of drug-resistant HIV variants, which can be transmitted to treatment-naive individuals, thereby complicating treatment strategies (Clutter et al., 2016). In low- and middle-income countries (LMICs), the scale-up of ART has similarly led to the emergence of drug-resistant strains, although the patterns and prevalence of resistance can differ due to variations in treatment practices and access to diagnostic testing (Figure 3) (Clutter et al., 2016).
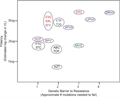 Figure 3 ARV potency versus genetic barrier to resistance (Adopted from Clutter et al., 2016) Image caption: ARV: antiretroviral; VL: viral load; ABC: abacavir; ATV/r: boosted atazanavir; DRV/r: boosted darunavir; DTG: dolutegravir; EFV: efavirenz; FTC: emtricitabine; EVG: elvitegravir; T20: enfuvirtide; ETR: etravirine; 3TC: lamivudine; LPV/r: boosted lopinavir; MVC: maraviroc; NVP: nevirapine; RAL: raltegravir; RPV: rilpivirine; and TDF: tenofovir. ARVs in black font are nucleoside or nucleotide reverse transcriptase inhibitors (NRTIs), those in purple font are non-NRTIs (NNRTIs), those in blue font are protease inhibitors (PIs), those in red font are integrase strand transfer inhibitors (INSTIs), and those in green font are entry inhibitors. ARVs appearing together in the same ellipse should be considered to have roughly equivalent potencies and genetic barriers to resistance (Adopted from Clutter et al., 2016) |
In regions with high HIV prevalence, such as sub-Saharan Africa, the emergence of MDR HIV is particularly concerning. The genetic diversity of HIV in these regions, coupled with the high mutation rate of the virus, facilitates the rapid development of resistance to multiple drugs (Martinez-Picado and Martínez, 2008). Studies have shown that the prevalence of transmitted drug resistance (TDR) and acquired drug resistance (ADR) is increasing, posing a significant challenge to the effectiveness of ART programs (Clutter et al., 2016). The World Health Organization (WHO) has recommended surveillance of pre-treatment drug resistance (PDR) to monitor and address this issue (Clutter et al., 2016).
6.2 Molecular Characterization of Resistance Mechanisms
The molecular mechanisms underlying HIV drug resistance are complex and involve multiple genetic mutations that confer resistance to different classes of antiretroviral drugs. Resistance to reverse transcriptase (RT) inhibitors, both nucleoside (NRTIs) and non-nucleoside (NNRTIs), is a major concern. These mutations can significantly impact the fitness of the virus, although compensatory mutations can restore replication capacity to some extent (Martinez-Picado and Martínez, 2008).
The genetic barrier to resistance, which refers to the difficulty for the virus to develop resistance mutations, varies among different drugs. For instance, the addition of lamivudine to zidovudine increases the genetic barrier to zidovudine resistance but decreases it for didanosine (Beerenwinkel et al., 2005). This highlights the importance of understanding the specific resistance pathways and the interactions between different drugs in designing effective treatment regimens.
Recent advances in bioinformatics have facilitated the study of resistance mechanisms. Machine learning approaches can predict the level of resistance from HIV sequence data, while phylogenetic methods can track the emergence and dynamics of resistant HIV transmission clusters. Deep sequencing technologies have further enhanced our understanding of within-host and between-host genetic diversity, providing insights into the evolution of resistance (Blassel et al., 2021).
6.3 Impact on Treatment Outcomes and Public Health Responses
The emergence of MDR HIV strains has significant implications for treatment outcomes and public health responses. Patients infected with drug-resistant HIV variants are at a higher risk of virological failure, which can lead to disease progression and increased mortality (Clutter et al., 2016). This necessitates the use of more complex and often more expensive treatment regimens, which may not be readily available in resource- limited settings.
Public health responses to MDR HIV must be multifaceted, involving both prevention and management strategies. Surveillance of drug resistance is crucial to inform treatment guidelines and ensure the effectiveness of ART programs. In high-income countries, routine resistance testing is part of standard care, allowing clinicians to tailor treatment regimens to individual patients and prevent the accumulation of further resistance. In LMICs, where access to diagnostic testing is limited, there is a need for innovative approaches to monitor and manage drug resistance (Clutter et al., 2016).
Mathematical models have been used to study the dynamics of drug resistance and evaluate the effectiveness of different treatment strategies. For example, improved gradual dosage reduction (iGDR) has been shown to decrease the threat of resistant virus emergence compared to structured treatment interruption (STI). These models can help in designing treatment protocols that minimize the risk of resistance development.
In conclusion, the emergence of MDR HIV strains poses a significant challenge to the global fight against HIV/AIDS. Understanding the molecular mechanisms of resistance, monitoring the prevalence of resistant strains, and implementing effective public health responses are essential to mitigate the impact of drug resistance on treatment outcomes and public health. Continued research and innovation are needed to stay ahead of the evolving virus and ensure the long-term success of ART programs.
7 Impact of HIV Adaptation on Vaccine Development
7.1 Challenges posed by HIV’s genetic diversity for vaccine design
The genetic diversity of HIV-1 is one of the most significant challenges in developing an effective vaccine. HIV-1 exhibits extraordinary variability, with envelope proteins differing by more than 30% in their amino acid sequences across different strains (Gaschen et al., 2002). This diversity is driven by the high mutation rate of the virus, facilitated by its error-prone reverse transcriptase, and the high rate of viral replication, which produces billions of new virions each day (Lal et al., 2005). The extensive glycosylation of the HIV envelope glycoprotein (gp120) further complicates vaccine design, as it shields the virus from neutralizing antibodies (Greenspan, 2014).
The diversity of HIV-1 is not uniform across all viral proteins, with the greatest variation observed in the envelope glycoproteins. This variation is a result of the virus's adaptation to host immune responses, leading to the evolution of escape mutants (Mcburney and Ross, 2008). The high level of diversity within and between clades of HIV-1 means that a vaccine effective against one clade may not be effective against another, posing a significant obstacle to the development of a universally effective vaccine (Nabel et al., 2002).
7.2 Overview of current vaccine candidates and their limitations
Current HIV vaccine candidates have shown limited success, primarily due to the challenges posed by the virus's genetic diversity. Most candidate vaccines are derived from specific isolates, with the hope that they will be cross-reactive enough to protect against a broad range of circulating viruses. However, this approach may be overly optimistic given the extent of HIV-1 diversity (Gaschen et al., 2002).
One of the most notable vaccine trials, RV144, demonstrated modest efficacy but also highlighted the issue of viral adaptation. Breakthrough infections in the trial were associated with HIV genotypes that had differential rates of transmission in vaccine and placebo recipients, suggesting that the virus can rapidly adapt to vaccine-induced immune pressures (Herbeck et al., 2018). This rapid adaptation can diminish the overall efficacy of the vaccine and reduce the number of infections averted by vaccination programs.
Another approach involves the use of centralized or polyvalent sequence designs to elicit broadly reactive immune responses. These designs aim to reduce the genetic distance between the vaccine strain and contemporary circulating viruses, but their effectiveness in eliciting long-lasting protective immunity remains to be fully demonstrated (Mcburney and Ross, 2008).
7.3 Innovative approaches to developing an effective HIV vaccine
To overcome the challenges posed by HIV-1's genetic diversity, several innovative approaches are being explored. One promising strategy involves the design of T cell immunogens that focus on the most conserved regions of the HIV-1 proteome. For example, the HIVCONSV vaccine candidate assembles the 14 most conserved regions of the HIV-1 proteome into a single chimeric protein, which has shown the ability to induce HIV-1-specific T cell responses in pre-clinical studies (Létourneau et al., 2007).
Another approach is the development of mosaic immunogens, which are designed to contain multiple potential T cell epitope variants. These immunogens aim to provide broad coverage of the diverse HIV-1 population by including sequences from different clades and subtypes (Tongo and Burgers, 2014). Early clinical studies have shown that these mosaic immunogens can elicit cross-reactive T cell responses, offering a potential pathway to a more universally effective vaccine.
Polyvalent vaccine strategies are also being evaluated, which involve the use of multiple antigens to improve the coverage of epitope diversity and enhance the cross-reactivity of immune responses. These strategies include the use of full proteins in novel vectors and concatenated conserved protein regions. Additionally, lineage-based design strategies are being explored to recapitulate and accelerate the natural development of broadly neutralizing antibodies (bNAbs) in vivo, inspired by longitudinal studies of bNAb emergence in HIV-1 infection (Korber et al., 2017).
In conclusion, while the genetic diversity of HIV-1 presents significant challenges for vaccine development, innovative approaches focusing on conserved regions, mosaic immunogens, and polyvalent strategies offer promising avenues for the development of an effective HIV vaccine. Continued research and international collaboration will be essential to advance these candidates into human testing and efficacy trials.
8 Host Factors Influencing HIV Adaptation
8.1 Role of Host Genetics in Susceptibility to HIV Infection
Host genetics play a pivotal role in determining susceptibility to HIV infection and the progression of the disease. One of the most significant genetic factors is the human leukocyte antigen (HLA) system. Specific HLA alleles, such as HLA-B27 and HLA-B57, have been associated with better control of HIV infection due to their ability to present viral epitopes effectively to CD8+T cells, facilitating a robust immune response (Kawashima et al., 2009; Martin and Carrington, 2013; Crux and Elahi, 2017). Conversely, mutations within these epitopes can allow the virus to escape immune detection, highlighting the dynamic interplay between host genetics and viral evolution (Kawashima et al., 2009).
Another critical genetic factor is the chemokine receptor CCR5. The CCR5-Δ32 mutation, which results in a non-functional receptor, provides resistance to HIV infection in homozygous individuals and delays disease progression in heterozygous individuals (Hogan and Hammer, 2001; Martin and Carrington, 2013). This mutation prevents the virus from using CCR5 as a co-receptor for entry into host cells, thereby impeding infection. However, HIV can adapt by using alternative co-receptors such as CXCR4, demonstrating the virus's ability to evolve in response to host genetic defenses (Hogan and Hammer, 2001).
Genome-wide association studies (GWAS) have further identified other genetic variants that influence HIV susceptibility and disease progression. These studies underscore the complexity of host genetic factors and their significant impact on the natural history of HIV infection (McLaren and Carrington, 2015; Tough and McLaren, 2019).
8.2 Impact of Co-Infections on HIV Evolution and Adaptation
Co-infections can significantly influence HIV evolution and adaptation. For instance, co-infection with other viruses such as hepatitis C virus (HCV) can modulate the immune response, potentially affecting HIV replication and disease progression (Crux and Elahi, 2017). The immune system's response to co-infections can create an environment that either suppresses or facilitates HIV replication, depending on the nature of the immune response elicited.
The presence of other pathogens can also lead to immune activation, which provides more target cells for HIV infection and increases viral replication rates. This heightened state of immune activation can accelerate the evolution of HIV by increasing the opportunities for the virus to mutate and adapt (Williamson, 2003; Bozek and Lengauer, 2010). Additionally, co-infections can alter the cytokine milieu, influencing the expression of chemokine receptors and other factors critical for HIV entry and replication (Hogan and Hammer, 2001).
8.3 Influence of Host Behavior and Immune Status on HIV Dynamics
Host behavior and immune status are crucial determinants of HIV dynamics. Behavioral factors such as adherence to antiretroviral therapy (ART), sexual practices, and substance use can significantly impact viral load and the potential for HIV transmission. Poor adherence to ART can lead to suboptimal drug levels, allowing the virus to develop resistance mutations and adapt to the therapeutic pressure (Martinez-Picado and Martínez, 2008).
The immune status of the host, particularly the functionality of CD4+T cells and the overall immune activation level, plays a vital role in HIV dynamics. A robust immune response can control viral replication and slow disease progression, while immune exhaustion and chronic activation can facilitate viral persistence and evolution (Kawashima et al., 2009; Martin and Carrington, 2013). The interaction between the host's immune system and the virus is a continuous battle, with HIV constantly adapting to evade immune detection and destruction (Williamson, 2003).
Furthermore, the genetic variability of the host's immune response genes, such as those encoding HLA and KIR (killer cell immunoglobulin-like receptors), can influence the effectiveness of the immune response against HIV. Variations in these genes can affect the presentation of viral antigens and the activation of natural killer (NK) cells, thereby impacting the control of HIV infection (Martin and Carrington, 2013; Tough and McLaren, 2019).
In summary, the adaptation strategies of HIV are profoundly influenced by host genetics, co-infections, and host behavior and immune status. Understanding these factors is essential for developing effective interventions and therapies to combat HIV and mitigate its impact on affected populations. The interplay between host and viral factors underscores the complexity of HIV pathogenesis and the ongoing challenge of controlling this elusive enemy.
9 Future Directions in HIV Research and Treatment
9.1 Advances in Gene Editing and Functional Cure Strategies
Gene editing has emerged as a promising avenue for achieving a functional cure for HIV. The advent of CRISPR-Cas9 technology has enabled precise modifications to the genome, offering the potential to excise integrated HIV proviruses from infected cells. This approach aims to eliminate the latent reservoir of HIV, which is a significant barrier to curing the infection. Recent studies have demonstrated the feasibility of using gene editing to target and disrupt the CCR5 co-receptor, which HIV uses to enter cells. By knocking out CCR5, researchers hope to render cells resistant to HIV infection. Additionally, gene editing can be employed to enhance the immune response by modifying immune cells to better recognize and attack HIV-infected cells. These strategies are still in the experimental stages, but they hold great promise for the future of HIV treatment and potentially achieving a functional cure (Burton and Hangartner, 2016; Frattari et al., 2023).
9.2 The Potential of Broadly Neutralizing Antibodies (bNAbs)
Broadly neutralizing antibodies (bNAbs) have shown significant potential in both the treatment and prevention of HIV. These antibodies can neutralize a wide range of HIV strains by targeting conserved regions of the virus's envelope protein. Recent clinical trials have explored the use of bNAbs as a therapeutic option, either alone or in combination with other antiretroviral drugs. The results have been promising, with combinations of bNAbs effectively suppressing viral load in many patients. Moreover, bNAbs have been shown to enhance the host immune response when administered during the early stages of infection, potentially leading to better control of the virus (Burton and Hangartner, 2016; Halper-Stromberg and Nussenzweig, 2016; Frattari et al., 2023). Future research is focused on improving the efficacy and durability of bNAb-based therapies, as well as exploring their role in HIV cure strategies, such as in combination with immune modulators or therapeutic vaccines (Frattari et al., 2023).
9.3 The Role of Global Collaboration in Combating HIV’s Adaptive Challenge
Global collaboration is crucial in the fight against HIV, given the virus's ability to rapidly adapt and develop resistance to treatments. International partnerships and collaborative research efforts have been instrumental in advancing our understanding of HIV and developing new treatment strategies. For instance, the identification and characterization of bNAbs have been made possible through global networks of researchers and clinicians who share data and resources. Collaborative efforts are also essential for conducting large-scale clinical trials and ensuring that new treatments are accessible to populations worldwide. Furthermore, global health initiatives and funding from organizations such as the World Health Organization (WHO) and the Joint United Nations Programme on HIV/AIDS (UNAIDS) play a vital role in supporting research and implementing effective HIV prevention and treatment programs. Continued global collaboration will be key to overcoming the adaptive challenges posed by HIV and ultimately achieving the goal of ending the HIV epidemic (Burton and Hangartner, 2016; Halper-Stromberg and Nussenzweig, 2016; Frattari et al., 2023).
In conclusion, the future of HIV research and treatment is promising, with several innovative strategies on the horizon. Advances in gene editing and functional cure strategies, the potential of broadly neutralizing antibodies, and the importance of global collaboration are all critical components in the ongoing battle against HIV. By leveraging these approaches, researchers and clinicians hope to develop more effective treatments, achieve functional cures, and ultimately eradicate HIV. The road ahead is challenging, but with continued innovation and collaboration, the elusive enemy that is HIV can be overcome.
10 Concluding Remarks
HIV's ability to adapt to various immune pressures, particularly those associated with human leukocyte antigen (HLA), has been a significant focus of recent research. Studies have shown that HLA is a major driver of HIV evolution at both individual and population levels, with the virus adapting to the immunogenetic profiles of different human ethnic groups. This adaptation has substantial clinical and immunologic consequences, including impaired immunogenicity and accelerated CD4+T cell decline when pre-adapted viruses are transmitted. Additionally, viral adaptation to HLA-II has been found to decrease CD4+T-cell responses in vaccine recipients, potentially impacting the efficacy of HIV vaccines. These findings underscore the complexity of HIV's adaptation mechanisms and the challenges they pose to prevention and cure strategies.
The insights gained from understanding HIV adaptation have several implications for future research and treatment approaches. Identifying immunogenic viral epitopes with high mutational barriers to escape can inform the design of more effective vaccines. Additionally, the concept of "personalized" immunogen selection based on HLA-driven adapted and nonadapted viral forms could enhance HIV cure strategies. The need for novel prevention technologies, such as multipurpose prevention technologies (MPTs), is also highlighted, given the limitations of existing strategies in reducing global HIV incidence. Furthermore, the adaptation of digital health interventions (DHIs) to new contexts and populations can accelerate the scale-up of effective prevention and treatment measures. Addressing implementation barriers at the provider and system levels, particularly for marginalized groups like men who have sex with men (MSM) and people who inject drugs (PWID), is crucial for the successful deployment of HIV-related health innovations.
The ongoing challenge of HIV adaptation necessitates a call to action for continued innovation in HIV prevention and management. Researchers and healthcare providers must collaborate to develop and implement strategies that account for the virus's ability to evade immune responses. This includes advancing vaccine research to target nonadapted epitopes and exploring the potential of personalized immunogen selection. Additionally, there is a need for sustained efforts to overcome structural barriers to HIV prevention, such as the criminalization of drug use, which hinder the implementation of effective interventions for PWID. Public and political advocacy will be essential to integrate innovative HIV prevention approaches with addiction science and create effective drug policies. By leveraging the latest scientific insights and fostering a collaborative approach, we can make significant strides toward eliminating HIV transmission and improving the lives of those affected by the virus.
![]() Acknowledgments
Acknowledgments
The EcoEvo Publiser extends sincere thanks to two anonymous peer reviewers for their feedback on the manuscript.
Conflict of Interest Disclosure
The authors affirm that this research was conducted without any commercial or financial relationships that could be construed as a potential conflict of interest.
Abram M., Ferris A., Das K., Quiñones O., Shao W., Tuske S., Alvord W., Arnold E., and Hughes S., 2014, Mutations in HIV-1 reverse transcriptase affect the errors made in a single cycle of viral replication, Journal of Virology, 88: 7589-7601.
https://doi.org/10.1128/JVI.00302-14
Ávila-Ríos S., Carlson J., John M., Mallal S., and Brumme Z., 2019, Clinical and evolutionary consequences of HIV adaptation to HLA: implications for vaccine and cure, Current Opinion in HIV and AIDS, 14: 194-204.
https://doi.org/10.1097/COH.0000000000000541
Azimi F., and Lee J., 2020, Structural perspectives on HIV-1 Vif and APOBEC3 restriction factor interactions, Protein Science, 29: 391-406.
https://doi.org/10.1002/pro.3729
Bayliss R., Wheeldon J., Caucheteux S., Niessen C., and Piguet V., 2020, Identification of host trafficking genes required for HIV-1 virological synapse formation in dendritic cells, Journal of Virology, 94(9): e01597-19.
https://doi.org/10.1128/JVI.01597-19
Beerenwinkel N., Däumer M., Sing T., Rahnenfuhrer J., Lengauer T., Selbig J., Hoffmann D., and Kaiser R., 2005, Estimating HIV evolutionary pathways and the genetic barrier to drug resistance, Journal of Infectious Diseases, 191(11): 1953-1960.
https://doi.org/10.1086/430005
Blassel L., Zhukova A., Villabona-Arenas C., Atkins K., Hué S., and Gascuel O., 2021, Drug resistance mutations in HIV: new bioinformatics approaches and challenges, Current Opinion in Virology, 51: 56-64.
https://doi.org/10.1016/j.coviro.2021.09.009
Bowen N., Oo A., and Kim B., 2022, Mechanistic interplay between HIV-1 reverse transcriptase enzyme kinetics and host SAMHD1 protein: viral myeloid-cell tropism and genomic mutagenesis, Viruses, 14(8): 1622.
https://doi.org/10.3390/v14081622
Bozek K., and Lengauer T., 2010, Positive selection of HIV host factors and the evolution of lentivirus genes, BMC Evolutionary Biology, 10: 186.
https://doi.org/10.1186/1471-2148-10-186
Burton D., and Hangartner L., 2016, Broadly neutralizing antibodies to HIV and their role in vaccine design, Annual Review of Immunology, 34: 635-659.
https://doi.org/10.1146/annurev-immunol-041015-055515
Cabrera-Rodríguez R., Pérez-Yanes S., Estévez-Herrera J., Márquez-Arce D., Cabrera C., Espert L., Blanco J., and Valenzuela-Fernández A., 2021, The interplay of HIV and autophagy in early infection, Frontiers in Microbiology, 12: 661446.
https://doi.org/10.3389/fmicb.2021.661446
Calado M., Pires D., Conceição C., Ferreira R., Santos-Costa Q., Anes E., and Azevedo-Pereira J., 2023, Cell-to-cell transmission of HIV-1 and HIV-2 from infected macrophages and dendritic cells to CD4+ T lymphocytes, Viruses, 15(5): 1030.
https://doi.org/10.3390/v15051030
Cameron P., Freudenthal P., Barker J., Gezelter S., Inaba K., and Steinman R., 1992, Dendritic cells exposed to human immunodeficiency virus type 1 transmit a vigorous cytopathic infection to CD4+ T cells, Science, 257(5068): 383-387.
https://doi.org/10.1126/science.1352913
Carlson J., Du V., Pfeifer N., Bansal A., Tan V., Power K., Brumme C., Kreimer A., DeZiel C., Fusi N., Schaefer M., Brockman M., Gilmour J., Price M., Kilembe W., Haubrich R., John M., Mallal S., Shapiro R., Frater J., Harrigan P., Ndung’u T., Allen S., Heckerman D., Sidney J., Allen T., Goulder P., Brumme Z., Hunter E., and Goepfert P., 2016, Impact of pre-adapted HIV transmission, Nature Medicine, 22: 606-613.
https://doi.org/10.1038/nm.4100
Choudhuri I., Biswas A., Haldane A., and Levy R., 2022, Contingency and entrenchment of drug-resistance mutations in HIV viral proteins, Journal of Physical Chemistry B, 126(50): 10622-10636.
https://doi.org/10.1021/acs.jpcb.2c06123
Clutter D., Jordan M., Bertagnolio S., and Shafer R., 2016, HIV-1 drug resistance and resistance testing, Infection, Genetics and Evolution, 46: 292-307.
https://doi.org/10.1016/j.meegid.2016.08.031
Crux N., and Elahi S., 2017, Human leukocyte antigen (HLA) and immune regulation: how do classical and non-classical HLA alleles modulate immune response to human immunodeficiency virus and hepatitis C virus infections?, Frontiers in Immunology, 8: 832.
https://doi.org/10.3389/fimmu.2017.00832
Cuevas J., Geller R., Garijo R., López-Aldeguer J., and Sanjuán R., 2015, Extremely high mutation rate of HIV-1 in vivo, PLoS Biology, 13(9): e1002251.
https://doi.org/10.1371/journal.pbio.1002251
Delannoy A., Poirier M., and Bell B., 2019, Cat and mouse: HIV transcription in latency, immune evasion and cure/remission strategies, Viruses, 11(3): 269.
https://doi.org/10.3390/v11030269
Frattari G., Caskey M., and Søgaard O., 2023, Broadly neutralizing antibodies for HIV treatment and cure approaches, Current Opinion in HIV and AIDS, 18: 157-163.
https://doi.org/10.1097/COH.0000000000000802
Gaschen B., Taylor J., Yusim K., Foley B., Gao F., Lang D., Novitsky V., Haynes B., Hahn B., Bhattacharya T., and Korber B., 2002, Diversity considerations in HIV-1 vaccine selection, Science, 296: 2354-2360.
https://doi.org/10.1126/science.1070441
Greenspan N., 2014, Design challenges for HIV-1 vaccines based on humoral immunity, Frontiers in Immunology, 5: 335.
https://doi.org/10.3389/fimmu.2014.00335
Halper-Stromberg A. and Nussenzweig M., 2016, Towards HIV-1 remission: potential roles for broadly neutralizing antibodies, The Journal of Clinical Investigation, 126(2): 415-423.
https://doi.org/10.1172/JCI80561
Herbeck J., Peebles K., Edlefsen P., Rolland M., Murphy J., Gottlieb G., Abernethy N., Mullins J., Mittler J., and Goodreau S., 2018, HIV population-level adaptation can rapidly diminish the impact of a partially effective vaccine, Vaccine, 36(4): 514-520.
https://doi.org/10.1016/j.vaccine.2017.12.004
Hogan C. and Hammer S., 2001, Host determinants in HIV infection and disease: Part 2: genetic factors and implications for antiretroviral therapeutics, Annals of Internal Medicine, 134: 978-996.
https://doi.org/10.7326/0003-4819-134-10-200105150-00012
Huang S., Ren Y., Thomas A., Chan D., Mueller S., Ward A., Patel S., Bollard C., Cruz C., Karandish S., Truong R., Macedo A., Bosque A., Kovacs C., Benko E., Piechocka-Trocha A., Wong H., Jeng E., Nixon D., Ho Y., Siliciano R., Walker B., and Jones R., 2018, Latent HIV reservoirs exhibit inherent resistance to elimination by CD8+ T cells, The Journal of Clinical Investigation, 128: 876-889.
https://doi.org/10.1172/JCI97555
Iyidogan P. and Anderson K., 2014, Current perspectives on HIV-1 antiretroviral drug resistance, Viruses, 6: 4095-4139.
https://doi.org/10.3390/v6104095
Kawashima Y., Pfafferott K., Frater J., Matthews P., Payne R., Addo M., Gatanaga H., Fujiwara M., Hachiya A., Koizumi H., Kuse N., Oka S., Duda A., Prendergast A., Crawford H., Leslie A., Brumme Z., Brumme C., Allen T., Brander C., Kaslow R., Tang J., Hunter E., Allen S., Mulenga J., Branch S., Roach T., John M., Mallal S., Ogwu A., Shapiro R., Prado J., Fidler S., Weber J., Pybus O., Klenerman P., Ndung’u T., Phillips R., Heckerman D., Harrigan P., Walker B., Takiguchi M., and Goulder P., 2009, Adaptation of HIV-1 to human leukocyte antigen class I, Nature, 458: 641-645.
https://doi.org/10.1038/nature07746
Korber B., Hraber P., Wagh K., and Hahn B., 2017, Polyvalent vaccine approaches to combat HIV-1 diversity, Immunological Reviews, 275: 230-244.
https://doi.org/10.1111/imr.12516
Lal R., Chakrabarti S., and Yang C., 2005, Impact of genetic diversity of HIV-1 on diagnosis, antiretroviral therapy and vaccine development, The Indian Journal of Medical Research, 121(4): 287-314.
Létourneau S., Im E., Mashishi T., Brereton C., Bridgeman A., Yang H., Dorrell L., Dong T., Korber B., McMichael A., and Hanke T., 2007, Design and pre-clinical evaluation of a universal HIV-1 vaccine, PLoS ONE, 2(10): e984.
https://doi.org/10.1371/journal.pone.0000984
Martin M. and Carrington M., 2013, Immunogenetics of HIV disease, Immunological Reviews, 254(1): 245-264.
https://doi.org/10.1111/imr.12071
Martinez-Picado J. and Martínez M., 2008, HIV-1 reverse transcriptase inhibitor resistance mutations and fitness: a view from the clinic and ex vivo, Virus Research, 134(1–2): 104-123.
https://doi.org/10.1016/j.virusres.2007.12.021
Martín-Moreno A. and Muñoz-Fernández M., 2019, Dendritic cells, the double agent in the war against HIV-1, Frontiers in Immunology, 10: 2485.
https://doi.org/10.3389/fimmu.2019.02485
McBurney S. and Ross T., 2008, Viral sequence diversity: challenges for AIDS vaccine designs, Expert Review of Vaccines, 7: 1405-1417.
https://doi.org/10.1586/14760584.7.9.1405
McDougal J., Nicholson J., Cross G., Cort S., Kennedy M., and Mawle A., 1986, Binding of the human retrovirus HTLV-III/LAV/ARV/HIV to the CD4 (T4) molecule: conformation dependence, epitope mapping, antibody inhibition, and potential for idiotypic mimicry, Journal of Immunology, 137(9): 2937-2944.
https://doi.org/10.4049/jimmunol.137.9.2937
McLaren P. and Carrington M., 2015, The impact of host genetic variation on infection with HIV-1, Nature Immunology, 16: 577-583.
https://doi.org/10.1038/ni.3147
Meriki H., Tufon K., Anong D., Atanga P., Anyangwe I., Cho-Ngwa F., and Nkuo-Akenji T., 2019, Genetic diversity and antiretroviral resistance-associated mutation profile of treated and naive HIV-1-infected patients from Cameroon, PLoS ONE, 14(11): e0225575.
https://doi.org/10.1371/journal.pone.0225575
Mujib S., Saiyed A., Fadel S., Bozorgzad A., Aidarus N., Yue F., Benko E., Kovacs C., Emert-Sedlak L., Smithgall T., and Ostrowski M., 2017, Pharmacologic HIV-1 Nef blockade promotes CD8 T cell-mediated elimination of latently HIV-1-infected cells in vitro, JCI Insight, 2(17): e93684.
https://doi.org/10.1172/jci.insight.93684
Mulherkar T., Gómez D., Sandel G., and Jain P., 2022, Co-infection and cancer: host–pathogen interaction between dendritic cells and HIV-1, HTLV-1, and other oncogenic viruses, Viruses, 14(9): 2037.
https://doi.org/10.3390/v14092037
Mylvaganam G., Yanez A., Maus M., and Walker B., 2019, Toward T cell-mediated control or elimination of HIV reservoirs: lessons from cancer immunology, Frontiers in Immunology, 10: 2109.
https://doi.org/10.3389/fimmu.2019.02109
Nabel G., Makgoba W., and Esparza J., 2002, HIV-1 diversity and vaccine development, Science, 296: 2335.
https://doi.org/10.1126/science.296.5577.2335
Paredes R. and Clotet B., 2010, Clinical management of HIV-1 resistance, Antiviral Research, 85(1): 245-265.
https://doi.org/10.1016/j.antiviral.2009.09.015
Poli G., 2013, Cell-to-cell vs. cell-free HIV-1 transmission from macrophages to CD4+ T lymphocytes: lessons from the virology textbook, AIDS, 27(14): 2307-2308.
https://doi.org/10.1097/QAD.0b013e328363619a
Shan L., Deng K., Gao H., Xing S., Capoferri A., Durand C., Rabi A., Laird G., Kim M., Hosmane N., Yang H., Zhang H., Margolick J., Li L., Cai W., Ke R., Flavell R., Siliciano J., and Siliciano R., 2017, Transcriptional reprogramming during effector-to-memory transition renders CD4+ T cells permissive for latent HIV-1 infection, Immunity, 47: 766-775.e3.
https://doi.org/10.1016/j.immuni.2017.09.014
Steiner M., Gibson K., and Crandall K., 2020, Drug resistance prediction using deep learning techniques on HIV-1 sequence data, Viruses, 12(5): 560.
https://doi.org/10.3390/v12050560
Ta T., Malik S., Anderson E., Jones A., Perchik J., Freylikh M., Sardo L., Klase Z., and Izumi T., 2022, Insights into persistent HIV-1 infection and functional cure: novel capabilities and strategies, Frontiers in Microbiology, 13: 862270.
https://doi.org/10.3389/fmicb.2022.862270
Tongo M. and Burgers W., 2014, Challenges in the design of a T cell vaccine in the context of HIV-1 diversity, Viruses, 6: 3968-3990.
https://doi.org/10.3390/v6103968
Tough R. and McLaren P., 2019, Interaction of the host and viral genome and their influence on HIV disease, Frontiers in Genetics, 9: 720.
https://doi.org/10.3389/fgene.2018.00720
Tsunetsugu-Yokota Y. and Muhsen M., 2013, Development of human dendritic cells and their role in HIV infection: antiviral immunity versus HIV transmission, Frontiers in Microbiology, 4: 178.
https://doi.org/10.3389/fmicb.2013.00178
Williamson S., 2003, Adaptation in the env gene of HIV-1 and evolutionary theories of disease progression, Molecular Biology and Evolution, 20(8): 1318-1325.
https://doi.org/10.1093/molbev/msg144
Yin X., Langer S., Zhang Z., Herbert K., Yoh S., König R., and Chanda S., 2020, Sensor sensibility—HIV-1 and the innate immune response, Cells, 9(1): 254.
https://doi.org/10.3390/cells9010254

. HTML
Associated material
. Readers' comments
Other articles by authors
. Caijuan Shou
. Jie Zhang
Related articles
. HIV adaptation
. Genetic diversity
. Antiretroviral resistance
. Immune escape
. Latent reservoir
Tools
. Post a comment


